We do not cover all features of the packages. Please refer to their documentation to learn them.
https://
github .com /apache /arrow -julia (Arrow.jl)
Here we’ll load CSV.jl to read and write CSV files and Arrow.jl and JLD2.jl, which allow us to work with a binary format, and finally JSONTables.jl for JSON interaction.
using DataFrames
using Arrow
using CSV
using JSONTables
using CodecZlib
using ZipFile
using JLD2
using StatsPlots ## for charts
using Mmap ## for zip compressionLet’s create a simple DataFrame for testing purposes,
x = DataFrame(
A=[true, false, true], B=[1, 2, missing],
C=[missing, "b", "c"], D=['a', missing, 'c']
)and use eltypes to look at the column-wise types.
eltype.(eachcol(x))4-element Vector{Type}:
Bool
Union{Missing, Int64}
Union{Missing, String}
Union{Missing, Char}CSV.jl¶
Let’s use CSV.jl to save x to disk; make sure x1.csv does not conflict with some file in your working directory.
tmpdir = mktempdir()
location = joinpath(tmpdir, "x1.csv")
CSV.write(location, x)"/tmp/jl_SCEErZ/x1.csv"Now we can see how it was saved by reading x.csv.
print(read(location, String))A,B,C,D
true,1,,a
false,2,b,
true,,c,c
We can also load the CSV file back as a dataframe
y = CSV.read(location, DataFrame)Note that when loading in a DataFrame from a CSV the column type for columns :C :D have changed to use special strings defined in the InlineStrings.jl package.
eltype.(eachcol(y))4-element Vector{Type}:
Bool
Union{Missing, Int64}
Union{Missing, InlineStrings.String1}
Union{Missing, InlineStrings.String1}JSONTables.jl¶
Often you might need to read and write data stored in JSON format. JSONTables.jl provides a way to process them in row-oriented or column-oriented layout. We present both options below.
location1 = joinpath(tmpdir, "x1.json")
open(io -> arraytable(io, x), location1, "w")106location2 = joinpath(tmpdir, "x2.json")
open(io -> objecttable(io, x), location2, "w")76Read them back.
print(read(location1, String))[{"A":true,"B":1,"C":null,"D":"a"},{"A":false,"B":2,"C":"b","D":null},{"A":true,"B":null,"C":"c","D":"c"}]print(read(location2, String)){"A":[true,false,true],"B":[1,2,null],"C":[null,"b","c"],"D":["a",null,"c"]}y1 = open(jsontable, location1) |> DataFrameeltype.(eachcol(y1))4-element Vector{Type}:
Bool
Union{Missing, Int64}
Union{Missing, String}
Union{Missing, String}y2 = open(jsontable, location2) |> DataFrameeltype.(eachcol(y2))4-element Vector{Type}:
Bool
Union{Missing, Int64}
Union{Missing, String}
Union{Missing, String}JLD2.jl¶
JLD2.jl is a high-performance, pure Julia library for saving and loading arbitrary Julia data structures, with HDF5 format.
Documentation: https://
save()andload(): General save and load using theFileIO.jlinterfacejldsave()andjldloac(): Advanced save and load with more optionssave_object()andload_object(): Single-object load and save
using JLD2
location = joinpath(tmpdir, "x.jld2")
save_object(location, x)Read it back.
load_object(location)Arrow.jl¶
Finally we use Apache Arrow format that allows, in particular, for data interchange with R or Python.
location = joinpath(tmpdir, "x.arrow")
Arrow.write(location, x)"/tmp/jl_SCEErZ/x.arrow"y = Arrow.Table(location) |> DataFrameeltype.(eachcol(y))4-element Vector{Type}:
Bool
Union{Missing, Int64}
Union{Missing, String}
Union{Missing, Char}Note that columns of y are immutable.
try
y.A[1] = false
catch e
show(e)
endReadOnlyMemoryError()This is because Arrow.Table uses memory mapping and thus uses a custom vector types:
y.A3-element Arrow.BoolVector{Bool}:
1
0
1y.B3-element Arrow.Primitive{Union{Missing, Int64}, Vector{Int64}}:
1
2
missingYou can get standard Julia Base vectors by copying a dataframe.
y2 = copy(y)y2.A3-element Vector{Bool}:
1
0
1y2.B3-element Vector{Union{Missing, Int64}}:
1
2
missingBasic benchmarking¶
Next, we’ll create some files in the temp directory.
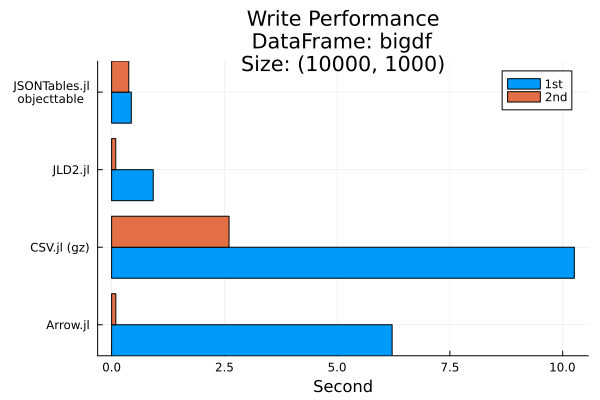
In particular, we’ll time how long it takes us to write a DataFrame with 1000 rows and 100000 columns.
bigdf = DataFrame(rand(Bool, 10^4, 1000), :auto)
bigdf[!, 1] = Int.(bigdf[!, 1])
bigdf[!, 2] = bigdf[!, 2] .+ 0.5
bigdf[!, 3] = string.(bigdf[!, 3], ", as string")
tmpdir = mktempdir()"/tmp/jl_S1zwiV"println("First run")First run
println("CSV.jl")
fname = joinpath(tmpdir, "bigdf1.csv.gz")
csvwrite1 = @elapsed @time CSV.write(fname, bigdf; compress=true)
println("Arrow.jl")
fname = joinpath(tmpdir, "bigdf.arrow")
arrowwrite1 = @elapsed @time Arrow.write(fname, bigdf)
println("JSONTables.jl arraytable")
fname = joinpath(tmpdir, "bigdf1.json")
jsontablesawrite1 = @elapsed @time open(io -> arraytable(io, bigdf), fname, "w")
println("JSONTables.jl objecttable")
fname = joinpath(tmpdir, "bigdf2.json")
jsontablesowrite1 = @elapsed @time open(io -> objecttable(io, bigdf), fname, "w")
println("JLD2.jl")
fname = joinpath(tmpdir, "bigdf.jld2")
jld2write1 = @elapsed @time save_object(fname, bigdf; compress = ZstdFilter())
println("Second run")
println("CSV.jl")
fname = joinpath(tmpdir, "bigdf1.csv.gz")
csvwrite2 = @elapsed @time CSV.write(fname, bigdf; compress=true)
println("Arrow.jl")
fname = joinpath(tmpdir, "bigdf.arrow")
arrowwrite2 = @elapsed @time Arrow.write(fname, bigdf)
println("JSONTables.jl arraytable")
fname = joinpath(tmpdir, "bigdf1.json")
jsontablesawrite2 = @elapsed @time open(io -> arraytable(io, bigdf), fname, "w")
println("JSONTables.jl objecttable")
fname = joinpath(tmpdir, "bigdf2.json")
jsontablesowrite2 = @elapsed @time open(io -> objecttable(io, bigdf), fname, "w")
println("JLD2.jl")
fname = joinpath(tmpdir, "bigdf.jld2")
jld2write2 = @elapsed @time save_object(fname, bigdf; compress = ZstdFilter());
groupedbar(
repeat(["CSV.jl (gz)", "Arrow.jl", "JSONTables.jl\nobjecttable", "JLD2.jl"],
inner=2),
[csvwrite1, csvwrite2, arrowwrite1, arrowwrite2, jsontablesowrite1, jsontablesowrite2, jld2write1, jld2write2],
group=repeat(["1st", "2nd"], outer=4),
ylab="Second",
title="Write Performance\nDataFrame: bigdf\nSize: $(size(bigdf))",
permute = (:x, :y)
)CSV.jl
10.174846 seconds (45.07 M allocations: 1.591 GiB, 1.65% gc time, 74.85% compilation time: <1% of which was recompilation)
Arrow.jl
6.218923 seconds (6.63 M allocations: 325.350 MiB, 0.27% gc time, 98.42% compilation time)
JSONTables.jl arraytable
11.780088 seconds (229.63 M allocations: 5.497 GiB, 16.55% gc time, 0.12% compilation time: <1% of which was recompilation)
JSONTables.jl objecttable
0.436314 seconds (106.17 k allocations: 309.450 MiB, 47.97% gc time, 18.22% compilation time)
JLD2.jl
0.922709 seconds (1.62 M allocations: 87.091 MiB, 89.74% compilation time: 11% of which was recompilation)
Second run
CSV.jl
2.603549 seconds (44.41 M allocations: 1.560 GiB, 5.98% gc time)
Arrow.jl
0.093268 seconds (80.86 k allocations: 5.164 MiB)
JSONTables.jl arraytable
11.986314 seconds (229.63 M allocations: 5.497 GiB, 16.52% gc time, 0.08% compilation time)
JSONTables.jl objecttable
0.380219 seconds (20.83 k allocations: 305.240 MiB, 54.08% gc time, 2.52% compilation time)
JLD2.jl
0.091727 seconds (134.50 k allocations: 13.779 MiB)
data_files = ["bigdf1.csv.gz", "bigdf.arrow", "bigdf1.json", "bigdf2.json", "bigdf.jld2"] .|> (f -> joinpath(tmpdir, f))
df = DataFrame(file=["CSV.jl (gz)", "Arrow.jl", "objecttable", "arraytable", "JLD2.jl"], size=getfield.(stat.(data_files), :size))
sort!(df, :size)
@df df plot(:file, :size / 1024^2, seriestype=:bar, title="Format File Size (MB)", label="Size", ylab="MB")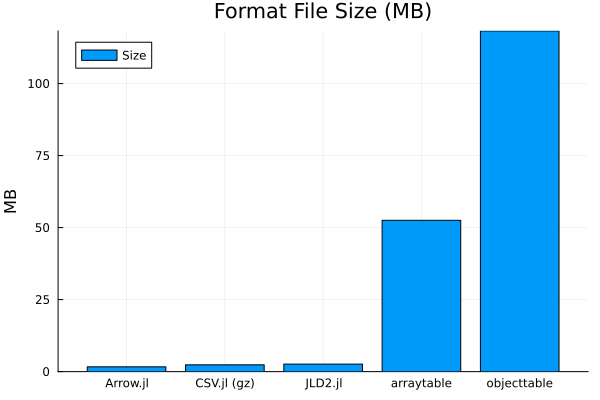
println("First run")
println("CSV.jl")
fname = joinpath(tmpdir, "bigdf1.csv.gz")
csvread1 = @elapsed @time CSV.read(fname, DataFrame)
println("Arrow.jl")
fname = joinpath(tmpdir, "bigdf.arrow")
arrowread1 = @elapsed @time df_tmp = Arrow.Table(fname) |> DataFrame
arrowread1copy = @elapsed @time copy(df_tmp)
println("JSONTables.jl arraytable")
fname = joinpath(tmpdir, "bigdf1.json")
jsontablesaread1 = @elapsed @time open(jsontable, fname)
println("JSONTables.jl objecttable")
fname = joinpath(tmpdir, "bigdf2.json")
jsontablesoread1 = @elapsed @time open(jsontable, fname)
println("JLD2.jl")
fname = joinpath(tmpdir, "bigdf.jld2")
jld2read1 = @elapsed @time load_object(fname)
println("Second run")
fname = joinpath(tmpdir, "bigdf1.csv.gz")
csvread2 = @elapsed @time CSV.read(fname, DataFrame)
println("Arrow.jl")
fname = joinpath(tmpdir, "bigdf.arrow")
arrowread2 = @elapsed @time df_tmp = Arrow.Table(fname) |> DataFrame
arrowread2copy = @elapsed @time copy(df_tmp)
println("JSONTables.jl arraytable")
fname = joinpath(tmpdir, "bigdf1.json")
jsontablesaread2 = @elapsed @time open(jsontable, fname)
println("JSONTables.jl objecttable")
fname = joinpath(tmpdir, "bigdf2.json")
jsontablesoread2 = @elapsed @time open(jsontable, fname)
println("JLD2.jl")
fname = joinpath(tmpdir, "bigdf.jld2")
jld2read2 = @elapsed @time load_object(fname);First run
CSV.jl
3.347728 seconds (4.26 M allocations: 216.272 MiB, 8.06% gc time, 29 lock conflicts, 113.16% compilation time)
Arrow.jl
0.530123 seconds (573.37 k allocations: 26.952 MiB, 98.61% compilation time)
0.053675 seconds (14.02 k allocations: 10.298 MiB)
JSONTables.jl arraytable
6.419493 seconds (271.15 k allocations: 1.772 GiB, 9.21% gc time, 0.06% compilation time)
JSONTables.jl objecttable
0.417452 seconds (7.39 k allocations: 566.946 MiB, 13.90% gc time, 0.03% compilation time)
JLD2.jl
0.309227 seconds (462.29 k allocations: 40.712 MiB, 87.92% compilation time)
Second run
1.248221 seconds (636.95 k allocations: 43.697 MiB, 17.87% gc time)
Arrow.jl
0.006109 seconds (84.07 k allocations: 3.593 MiB)
0.051327 seconds (14.02 k allocations: 10.298 MiB)
JSONTables.jl arraytable
6.532852 seconds (271.10 k allocations: 1.772 GiB, 9.32% gc time)
JSONTables.jl objecttable
0.406086 seconds (7.08 k allocations: 566.930 MiB, 10.23% gc time)
JLD2.jl
0.036153 seconds (121.76 k allocations: 24.191 MiB)
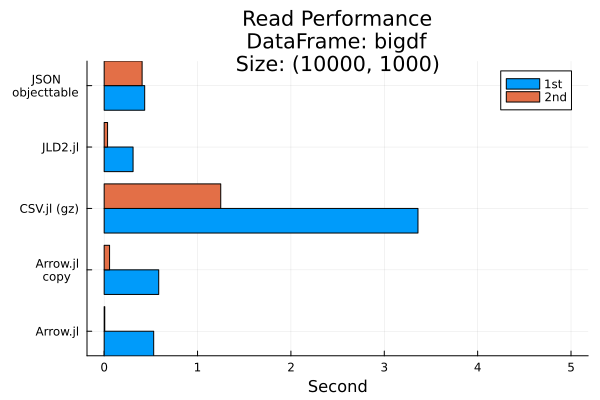
Exclude JSONTables due to much longer timing
groupedbar(
repeat(["CSV.jl (gz)", "Arrow.jl", "Arrow.jl\ncopy", ##"JSON\narraytable",
"JSON\nobjecttable", "JLD2.jl"], inner=2),
[csvread1, csvread2, arrowread1, arrowread2, arrowread1 + arrowread1copy, arrowread2 + arrowread2copy,
# jsontablesaread1, jsontablesaread2,
jsontablesoread1, jsontablesoread2, jld2read1, jld2read2],
group=repeat(["1st", "2nd"], outer=5),
ylab="Second",
title="Read Performance\nDataFrame: bigdf\nSize: $(size(bigdf))",
permute = (:x, :y)
)
Using gzip compression¶
A common user requirement is to be able to load and save CSV that are compressed using gzip. Below we show how this can be accomplished using CodecZlib.jl.
Again make sure that you do not have file named df_compress_test.csv.gz in your working directory.
We first generate a random data frame.
df = DataFrame(rand(1:10, 10, 1000), :auto)Use compress=true option to compress the CSV with the gz format.
tmpdir = mktempdir()
fname = joinpath(tmpdir, "df_compress_test.csv.gz")
CSV.write(fname, df; compress=true)"/tmp/jl_pXTEB8/df_compress_test.csv.gz"Read the CSV file back.
df2 = CSV.File(fname) |> DataFramedf == df2trueWorking with zip files¶
Sometimes you may have files compressed inside a zip file.
In such a situation you may use ZipFile.jl in conjunction an an appropriate reader to read the files.
Here we first create a ZIP file and then read back its contents into a DataFrame.
df1 = DataFrame(rand(1:10, 3, 4), :auto)df2 = DataFrame(rand(1:10, 3, 4), :auto)And we show yet another way to write a DataFrame into a CSV file:
Writing a CSV file into the zip file
w = ZipFile.Writer(joinpath(tmpdir, "x.zip"))
f1 = ZipFile.addfile(w, "x1.csv")
write(f1, sprint(show, "text/csv", df1))45write a second CSV file into the zip file
f2 = ZipFile.addfile(w, "x2.csv", method=ZipFile.Deflate)
write(f2, sprint(show, "text/csv", df2))46close(w)Now we read the compressed CSV file we have written:
r = ZipFile.Reader(joinpath(tmpdir, "x.zip"))
# find the index index of file called x1.csv
index_xcsv = findfirst(x -> x.name == "x1.csv", r.files)
# to read the x1.csv file in the zip file
df1_2 = CSV.read(read(r.files[index_xcsv]), DataFrame)df1_2 == df1true# find the index index of file called x2.csv
index_xcsv = findfirst(x -> x.name == "x2.csv", r.files)
# to read the x2.csv file in the zip file
df2_2 = CSV.read(read(r.files[index_xcsv]), DataFrame)df2_2 == df2trueNote that once you read a given file from r object its stream is all used-up (reaching its end). Therefore to read it again you need to close the file object r and open it again.
Also do not forget to close the zip file once you are done.
close(r)This notebook was generated using Literate.jl.