Sources:
using Plots
using Random
Random.seed!(2024)Random.TaskLocalRNG()Ticks size and properties¶
plot(sin, 0, 2π;
xticks=0:0.5:2π,
xrotation=60,
xtickfontsize=25,
bottom_margin=15Plots.mm)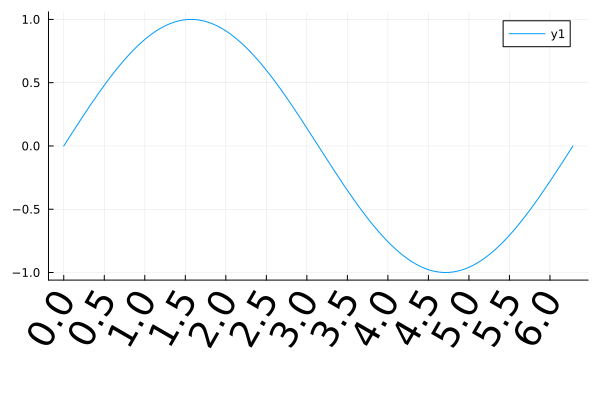
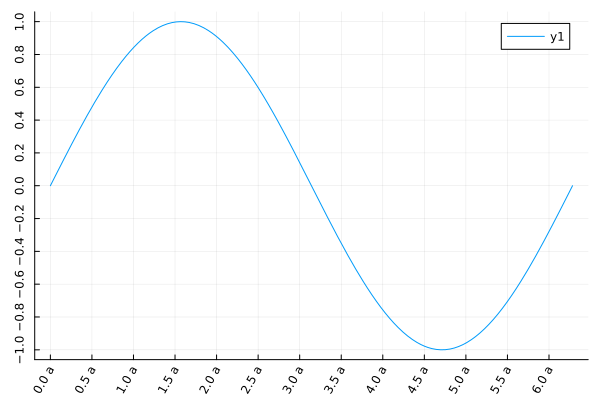
plot(sin, 0, 2π;
xtick=(0:0.5:2π, ["$i a" for i in 0:0.5:2π]),
ytick=-1:0.2:1,
xrotation=60,
yrotation=90,
)
No axis¶
axis=false
plot(sin, 0, 2π, axis=false)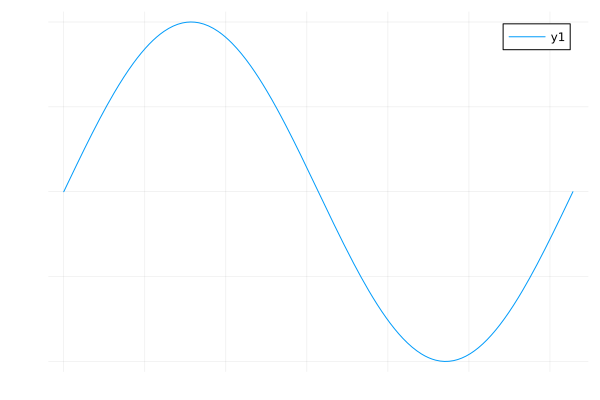
Log scale for axes¶
xscale=:log10, yscale=:log10
plot(exp, -5, 5, yscale=:log10, title="semilogy", legend=nothing)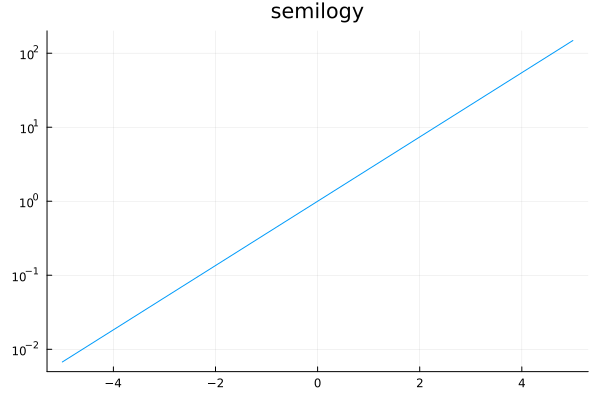
plot(log, 1e-5, 10, xscale=:log10, title="semilogx", legend=nothing)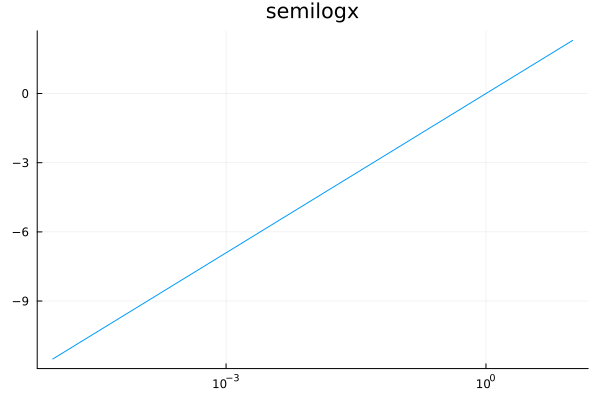
plot(x->x^1.7, 1e-3, 3, scale=:log10, title="log-log", label="x^(1.7)", legend=:topleft)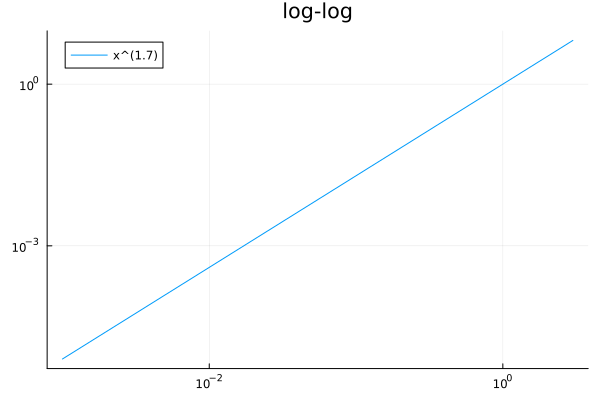
Axis range¶
xlims and ylims
plot(sin, 0, 2π, xlims=(-10, 10), ylims=(-2,2))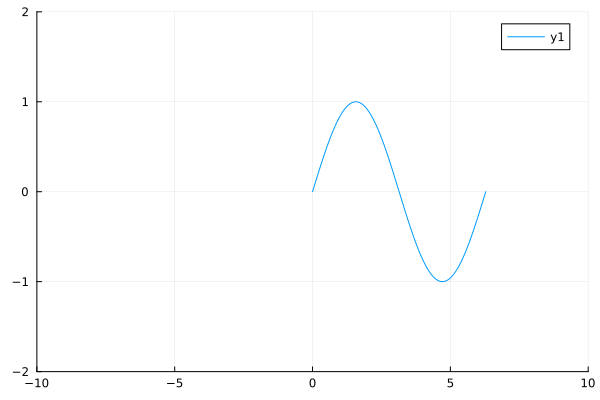
Scientific notation¶
yformatter=:scientific
plot(exp, 0, 10, yformatter=:scientific)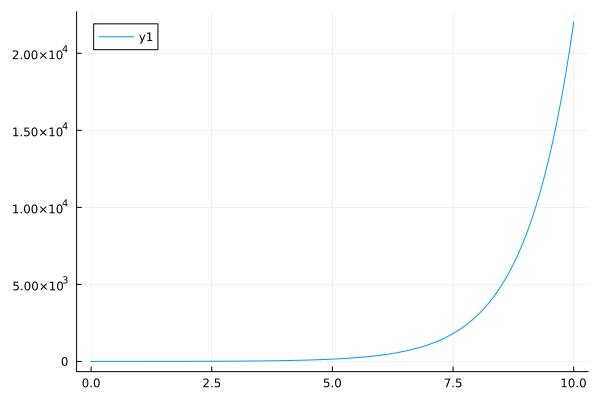
Flip¶
xflip=true and/or yflip=true
plot(identity, 0:0.01:2π, proj=:polar, xflip=true, yflip=true, legend=false)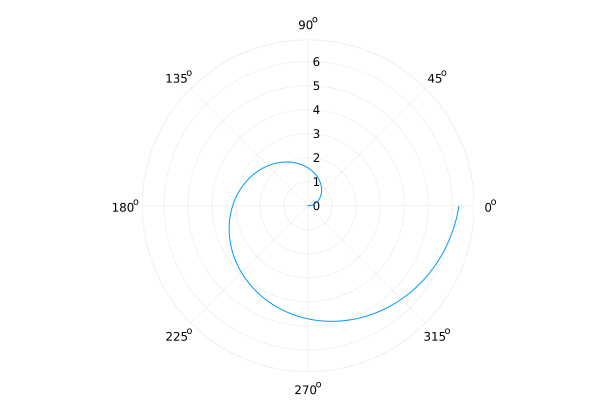
Aspect ratio¶
aspect_ratio=:equal or aspect_ratio=<number>
heatmap(bitrand(10, 10), aspect_ratio=:equal, c=:blues, colorbar=false)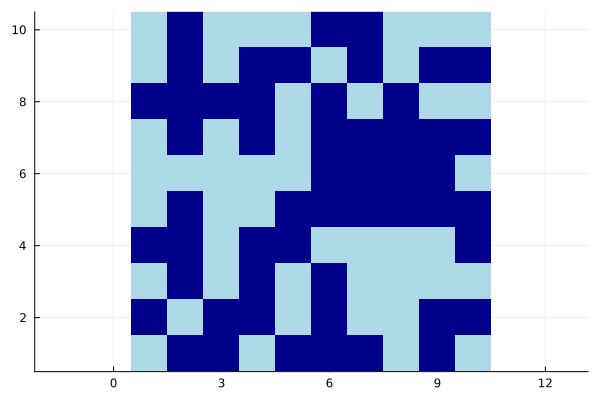
Fonts¶
LaTeX fonts are supported by the LaTeXStrings.jl package.
using Plots
using LaTeXStrings
plot(sin, 0, 2π,
title=L"y = \sin(x)",
titlefont=font(40), ## title
xlabel=L"x",
ylabel="y",
xguidefontsize=30, ## x-guide
yguidefontsize=20, ## y-guide
# guidefontsize=20, ## both x,y-guide
xtick=(0:0.5:2π, ["\$ $(i) \$" for i in 0:0.5:2π]),
ytick=-1:0.5:1,
xtickfontsize=15,
ytickfontsize=20,
# tickfontsize=10, ## for both x and y
label="Sin function",
legendfontsize=12,
xlims=(0,2π),
ylims=(-1,1),
bottom_margin=5Plots.mm,
left_margin=10Plots.mm,
top_margin=15Plots.mm
)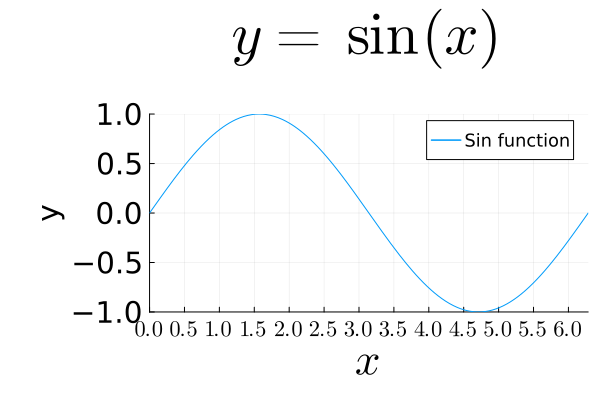
fib(x) = (((1+sqrt(5))/2)^x - ((1-sqrt(5))/2)^x)/sqrt(5)
ann = L"F_n = \frac{1}{\sqrt{5}} \left[\left( \frac{1+\sqrt{5}}{2} \right)^n - \left( \frac{1-\sqrt{5}}{2} \right)^n \right]"
plot(fib, 1:12, marker=:circle, xlabel=L"n", ylabel=L"F_n", annotation=(5, 100, ann))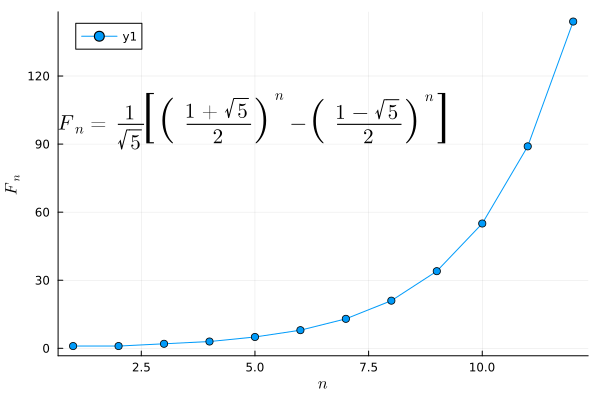
Bar plots¶
using Plots
using StatsPlots
using StatsBasemeasles = [38556, 24472, 14556, 18060, 19549, 8122, 28541, 7880, 3283, 4135, 7953, 1884]
mumps = [20178, 23536, 34561, 37395, 36072, 32237, 18597, 9408, 6005, 6268, 8963, 13882]
chickenPox = [37140, 32169, 37533, 39103, 33244, 23269, 16737, 5411, 3435, 6052, 12825, 23332]
ticklabel = string.(collect('A':'L'))12-element Vector{String}:
"A"
"B"
"C"
"D"
"E"
"F"
"G"
"H"
"I"
"J"
"K"
"L"Grouped vertical bar plots¶
Requires the StatsPlots.jl package.
groupedbar(data, bar_position = :dodge)
groupedbar([measles mumps chickenPox], bar_position = :dodge, bar_width=0.7, xticks=(1:12, ticklabel), label=["measles" "mumps" "chickenPox"])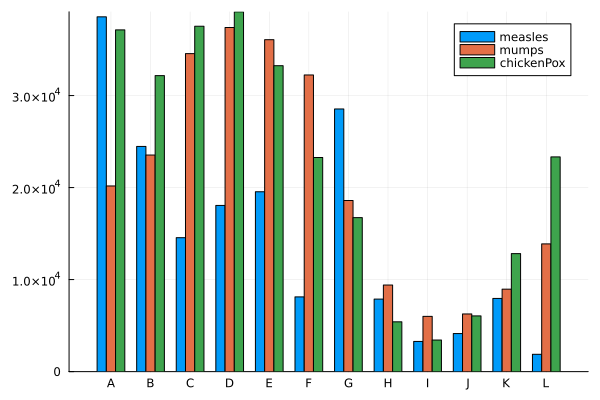
Stacked vertical bar plots¶
Requires StatsPlots package.
groupedbar(data, bar_position = :stack)
groupedbar([measles mumps chickenPox],
bar_position = :stack,
bar_width=0.7,
xticks=(1:12, ticklabel),
label=["measles" "mumps" "chickenPox"])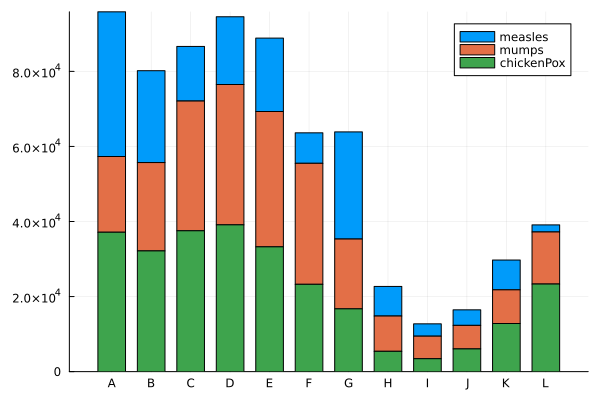
Horizontal Bar Plot¶
bar(data, orientation=:h)
bar(1:12, orientation=:h, yticks=(1:12, ticklabel), yflip=true)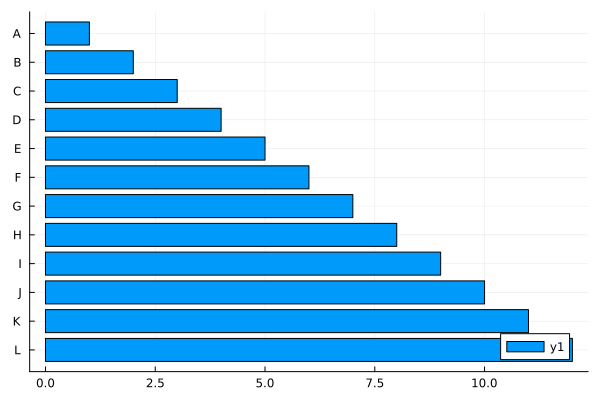
Categorical Histogram Plot¶
status = ["Poor", "Fair", "Good", "Excellent"]
data = sample(status, Weights([1,1,2,2]), 100)
datamap = countmap(data)
bar((x -> datamap[x]).(status), xticks=(1:4, status), legend=nothing)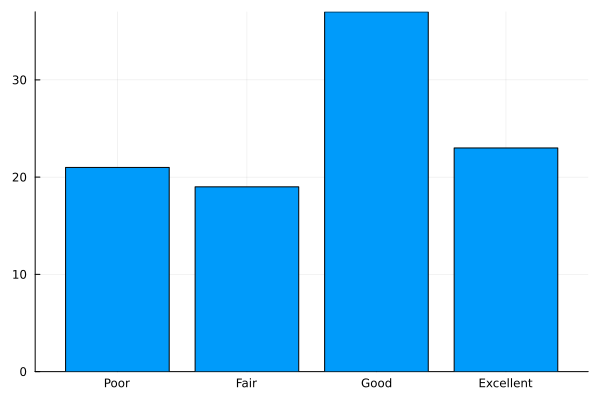
Histogram¶
histogram(data, bins=N)
using Plots
x = randn(1000)
y = randn(1000)
z = randn(1000)
histogram(x, bins=20, alpha=0.4, label="A")
histogram!(y, bins=20, alpha=0.4, label="B")
histogram!(z, bins=20, alpha=0.4, label="C")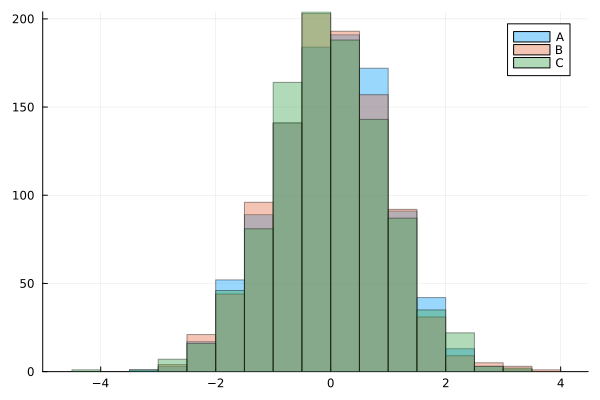
Box plots¶
using Plots
using StatsPlots
using Statistics
n = 30
science = rand(1:10, n)
boxplot(science, label="science")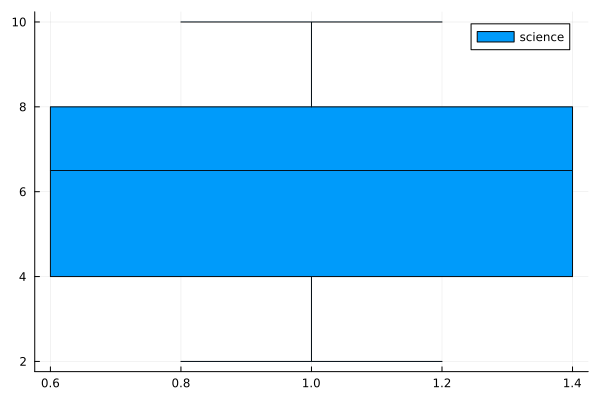
english = rand(1:10, n)
boxplot([science english], label=["science" "english"])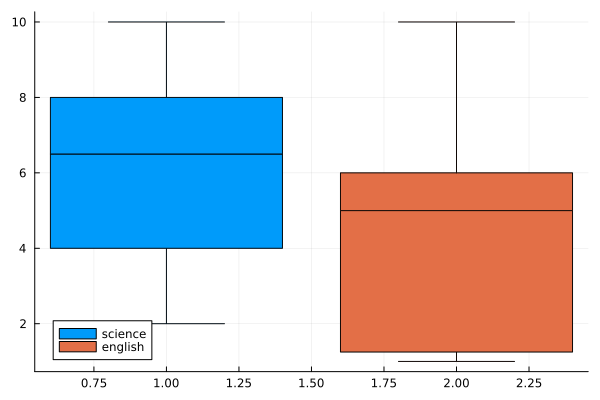
using Plots
xs = range(0, stop=2, length=50)
ys = range(0, stop=2, length=50)
f = (x , y) -> x^2 + y^2
contour(xs, ys, f)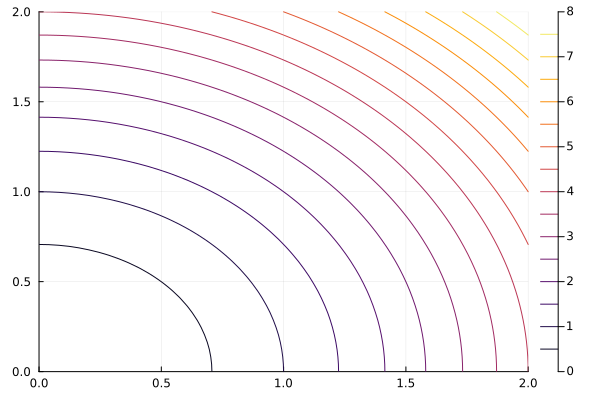
Nullclines¶
Nullclines (zero-growth isoclines) are curves where the derivative of one variable is zero. Nullclines are used to analyze evolution and stability of ODE systems.
using Plots
dx = (x, y) -> 3x - 1.4x*y
dy = (x, y) -> -y + 0.8x*y
myrange = -1:0.01:4
contour(myrange, myrange, dx, levels=[0], color=:red, legend=false)
contour!(myrange, myrange, dy, levels=[0], color=:blue, legend=false)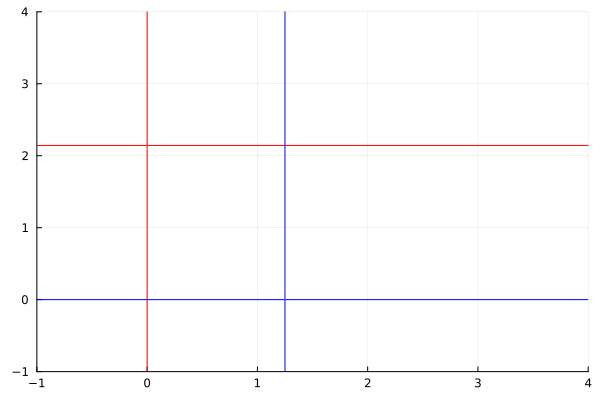
Contour plot over an array¶
contour(x1d, y1d, xy2d)
# Notice xs is transposed
# This makes zz a 2D matrix
zz = f.(xs', ys)
contour(xs, ys, zz)
Filled Contour Plots¶
contour(xs, ys, f, fill=true)contourf(xs, ys, f)
contour(0:0.01:5, 0:0.01:5, (x, y) -> sin(3x) * cos(x+y), xlabel="x", ylabel="y", fill=true)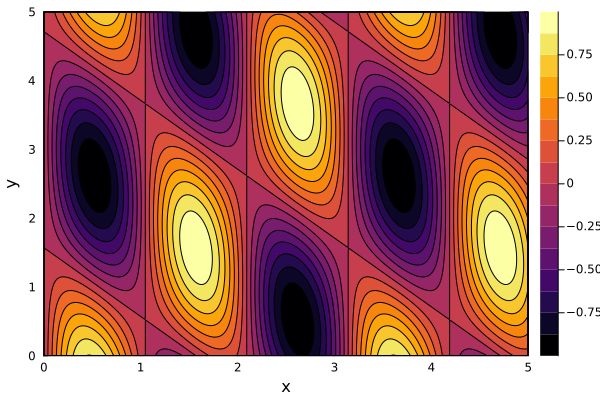
Datetime plot¶
Use
Datespackage andDatadata typeCustomize ticks
using Plots
using Dates
days = 31
position = cumsum(randn(days))
x = Date(2018,1,1):Day(1):Date(2018,1,31)
ticks = [x[i] for i in 1:5:length(x)]
plot(x, position,
xlabel="Date",
ylabel="Position",
title="Track of random walker",
legend=nothing,
xticks=ticks,
xrotation=45,
bottom_margin=10Plots.mm,
left_margin=5Plots.mm)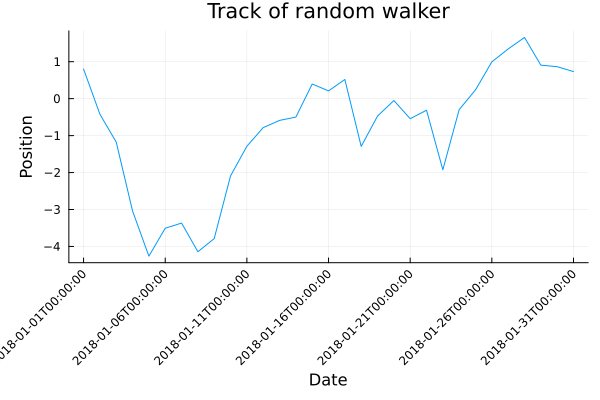
Error bar¶
plots(..., xerr=xerr, yerr=yerr)
using Plots
f = x -> 2 * x + 1
x = 0:0.1:2
n = length(x)
y = f.(x) + randn(n)
plot(x, y,
xerr=0.1 * rand(n),
yerr=rand(n),
legend=nothing)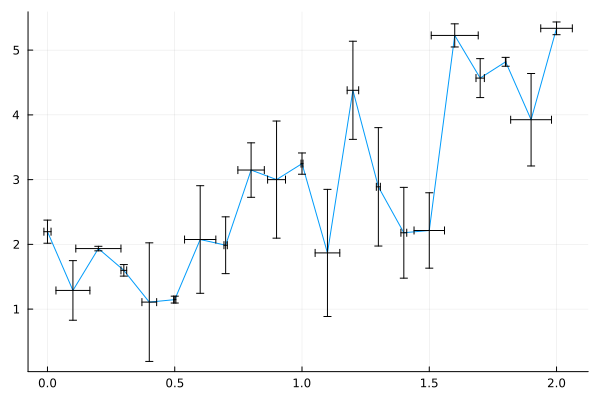
Heatmap¶
heatmap(data)
using Plots
a = rand(5,5)
xlabel = string.(collect('A':'E'))
ylabel = string.(collect('a':'e'))
heatmap(a, xticks=(1:5, xlabel),
yticks=(1:5, ylabel),
aspect_ratio=:equal)
fontsize = 15
nrow, ncol = size(a)
# Add number annotations to plots
ann = [(i,j, text(round(a[i,j], digits=2), fontsize, :white, :center))
for i in 1:nrow for j in 1:ncol]
annotate!(ann, linecolor=:white)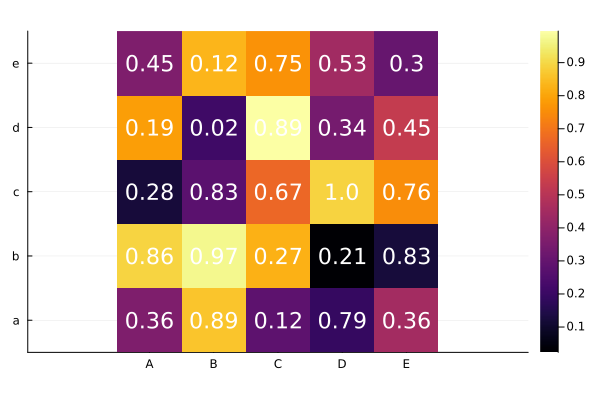
Line plots¶
using Plots
plot(x, y)
plot(f, xRange)
plot(f, xMin, xMax)
plot(x, [y1 y2])using Plots
# Data
x = 0:0.1:2pi
y1 = cos.(x)
y2 = sin.(x)
# Creating a plot in steps
plot(x, y1, color="blue", linewidth=3)
plot!(x, y2, color="red", line=:dash)
title!("Trigonometric functions")
xlabel!("angle")
ylabel!("sin(x) and cos(x)")
plot!(xlims=(0,2pi), ylims=(-2, 2), size=(600, 600))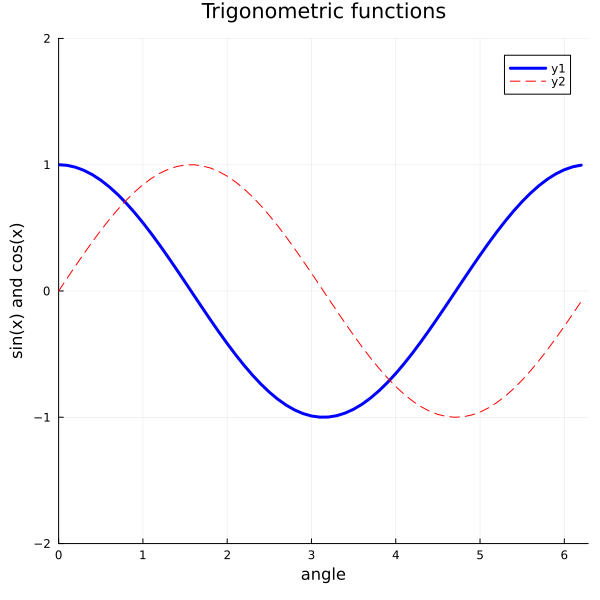
plot(x, y1, line=(:blue, 3))
plot!(x, y2, line=(:dash, :red))
# Use keywords to set the options all in one plot() call
plot!(title="Trigonometric functions",
xlabel="angle",
ylabel="sin(x) and cos(x)",
xlims=(0,2pi), ylims=(-2, 2), size=(600, 600))
Plotting multiple series¶
One row = one observation
One column = one species
time = 30
walker1 = cumsum(randn(time))
walker2 = cumsum(randn(time))
walker3 = cumsum(randn(time))
walker4 = cumsum(randn(time))
walker5 = cumsum(randn(time))
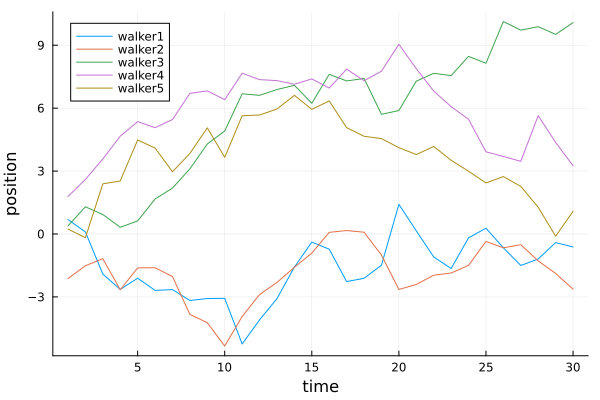
plot(1:time, [walker1 walker2 walker3 walker4 walker5],
xlabel="time", ylabel="position",
label=["walker1" "walker2" "walker3" "walker4" "walker5"],
legend=:topleft
)
Parametric plots¶
Functions can be plotted directly.
plot(f, xmin, xmax)plot(f, range_of_x)plot(fx(t), fy(t), range_of_t)
let
f = x -> 5exp(-x^2)
g = x -> x^2
plot([f, g], -3, 3, label=["f" "g"], legend=:top)
end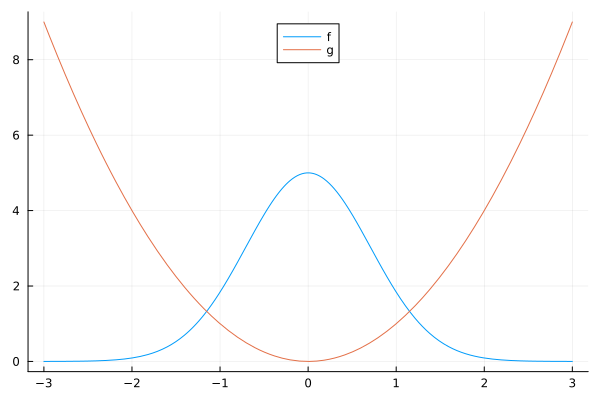
plot(sin, t->sin(2t), 0, 2π, leg=false, fill=(0,:orange))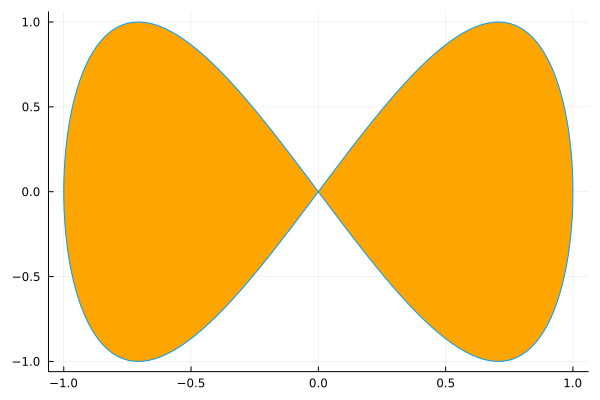
plot(cos, sin, t -> sin(5t), 0, 2pi, legend=nothing)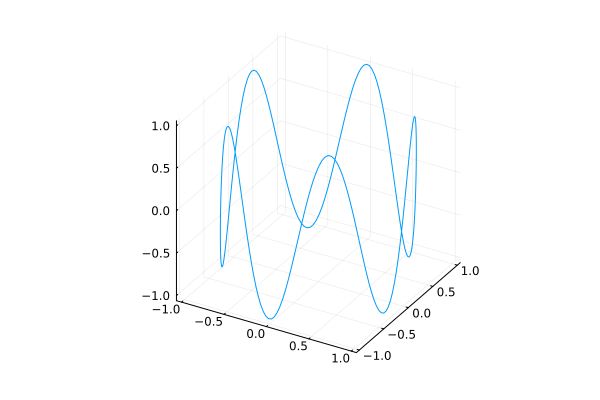
Line colors¶
plot(x, y, c=color)
using Plots
using SpecialFunctions
x = 0:0.2:10
y0 = besselj.(0,x)
y1 = besselj.(1,x)
y2 = besselj.(2,x)
y3 = besselj.(3,x)
y4 = besselj.(4,x)
y5 = besselj.(5,x)
y6 = besselj.(6,x)
colors = [:red :green :blue :cyan :magenta :yellow :black]
plot(x, [y0 y1 y2 y3 y4 y5 y6], c=colors)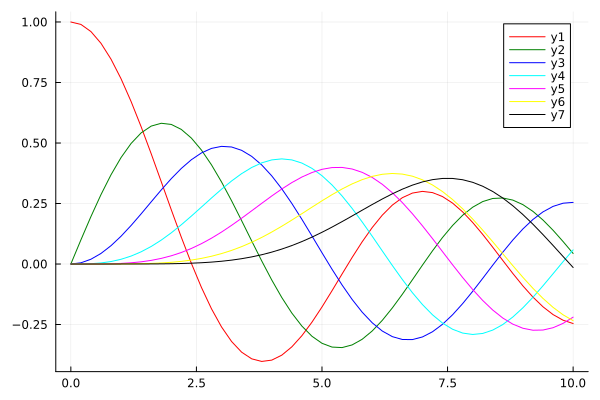
Line styles¶
using Plots
@show Plots.supported_markers();
@show Plots.supported_styles();Plots.supported_markers() = [:+, :auto, :circle, :cross, :diamond, :dtriangle, :heptagon, :hexagon, :hline, :ltriangle, :none, :octagon, :pentagon, :pixel, :rect, :rtriangle, :star4, :star5, :star6, :star7, :star8, :utriangle, :vline, :x, :xcross]
Plots.supported_styles() = [:auto, :dash, :dashdot, :dashdotdot, :dot, :solid]
style = Plots.supported_styles()[2:end]
style = reshape(style, 1, length(style))
plot(x, [y0 y1 y2 y3 y4], line=(3, style))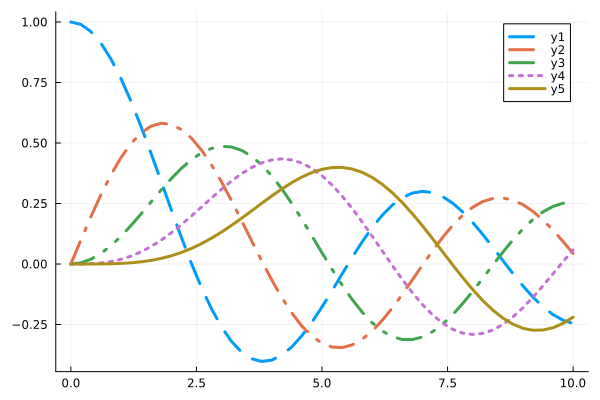
Polar Plots¶
plot(θ, r, proj=:polar)
using Plots
plot(θ -> 1 + cos(θ) * sin(θ)^2, 0, 2π, proj=:polar, lims=(0, 1.5))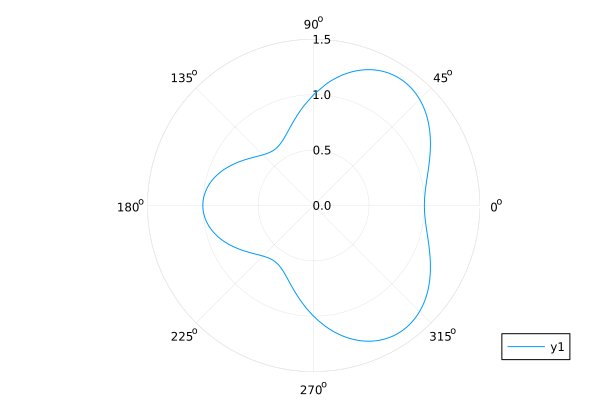
Rose Plots¶
plot(..., proj=:polar, line=:steppre)
n = 24
R = rand(n+1)
plot(0:2pi/n:2pi, R, proj=:polar, line=:steppre, lims=(0, 1), legend=nothing)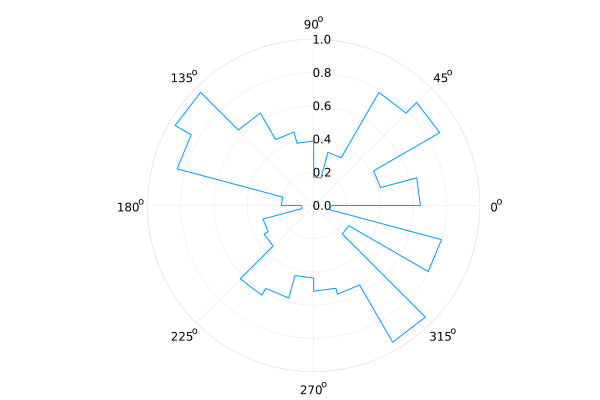
Quiver Plots¶
quiver(x1d, y1d, quiver=(vx1d, vy1d)quiver(x2d, y2d, quiver=(x, y)->(u, v))
using Plots
n = 7
f = (x,y) -> hypot(x, y) |> inv
x = repeat(-3:(2*3)/n:3, 1, n) |> vec
y = repeat(-3:(2*3)/n:3, 1, n)' |> vec
vx = f.(x,y) .* cos.(atan.(y,x)) |> vec
vy = f.(x,y) .* sin.(atan.(y,x)) |> vec
quiver(x, y, quiver=(vx, vy), aspect_ratio=:equal)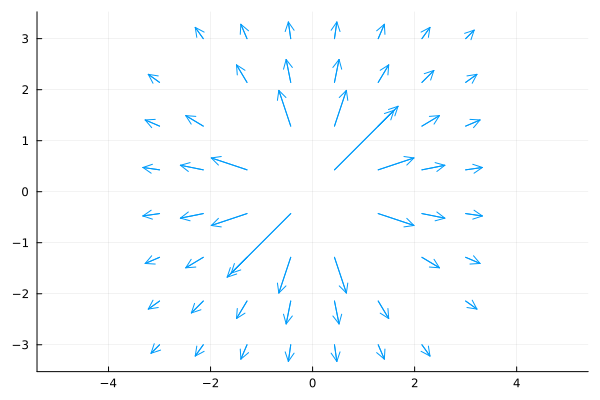
g = (x, y) -> [f(x,y) * cos(atan(y,x)), f(x,y) * sin(atan(y,x))]
xx = [x for y in -3:(2*3)/n:3, x in -3:(2*3)/n:3]
yy = [y for y in -3:(2*3)/n:3, x in -3:(2*3)/n:3]
quiver(xx, yy, quiver=g, aspect_ratio=:equal, color=:black)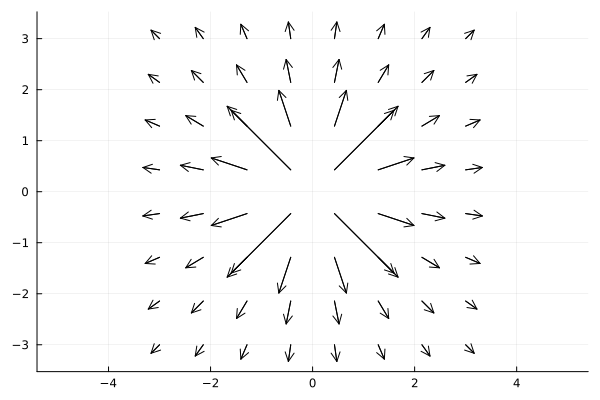
Scatter Plots¶
2D Scatter Plots: scatter(xpos, ypos)
using Plots
n = 50
x = rand(n)
y = rand(n)
ms = rand(50) * 30
scatter(x, y, markersize=ms)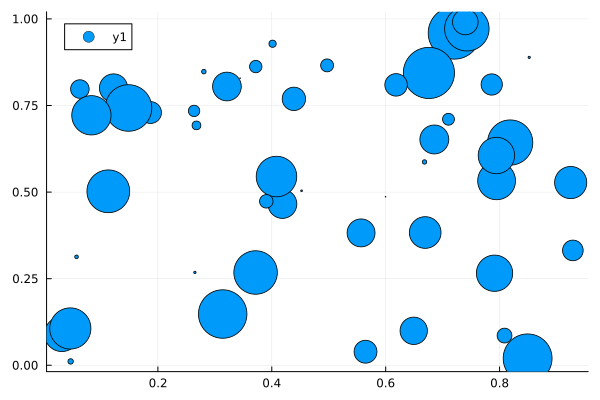
3D Scatter Plots: scatter(xpos, ypos, zpos)
scatter(x, y, rand(n), markersize=ms)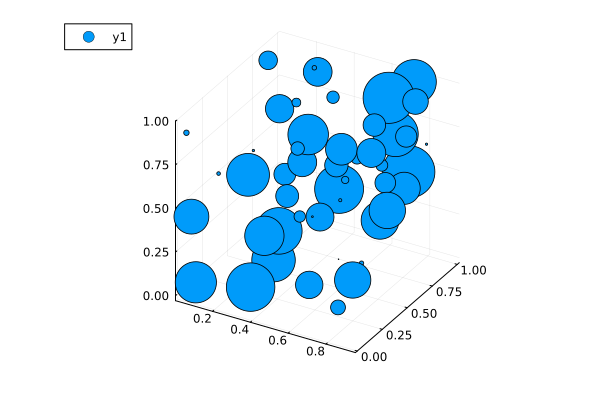
Shared Color bar¶
The trick is to
make an blank scatter plot for the colorbar.
use a dedicated space in the layout for the colorbar.
using Plots
x1 = range(0.0, 1.0, length=51)
y1 = range(-1.0, 0.0, length=51)
z1 = sin.(4*pi.*x1)*cos.(6*pi.*reshape(y1, 1, :))
x2 = range(0.25, 1.25, length=51)
y2 = range(-1.0, 0.0, length=51)
z2 = 2.0.*sin.(4*pi.*x2)*cos.(6*pi.*reshape(y2, 1, :))
x3 = range(0.0, 1.0, length=51)
y3 = range(-1.25, -0.25, length=51)
z3 = 3.0*sin.(4*pi.*x3)*cos.(6*pi.*reshape(y3, 1, :))
x4 = range(0.25, 1.25, length=51)
y4 = range(-1.25, -0.25, length=51)
z4 = 4.0.*sin.(4*pi.*x4)*cos.(6*pi.*reshape(y4, 1, :))
clims = extrema([z1; z2; z3; z4])
p1 = contourf(x1, y1, z1, clims=clims, c=:viridis, colorbar=false)
p2 = contourf(x2, y2, z2, clims=clims, c=:viridis, colorbar=false)
p3 = contourf(x3, y3, z3, clims=clims, c=:viridis, colorbar=false)
p4 = contourf(x4, y4, z4, clims=clims, c=:viridis, colorbar=false)
h2 = scatter(
[0,0], [0,1], zcolor=[0,3], clims=clims,
xlims=(1,1.1), label="", c=:viridis,
colorbar_title="cbar", framestyle=:none)
l = @layout [grid(2, 2) a{0.035w}]
p_all = plot(p1, p2, p3, p4, h2, layout=l, link=:all)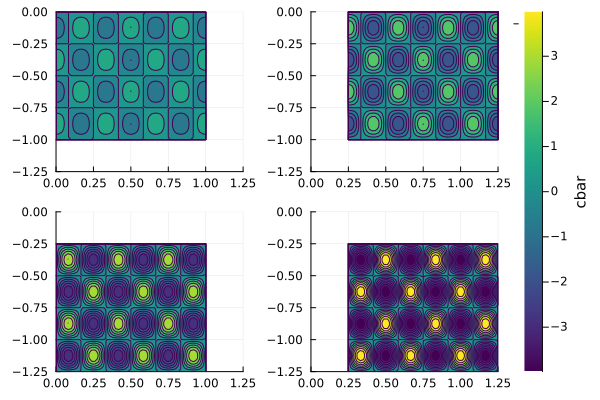
Stairstep plot¶
plot(..., line=:steppre)
plot(sin.(0:0.3:2pi), line=:steppre, label="Steps")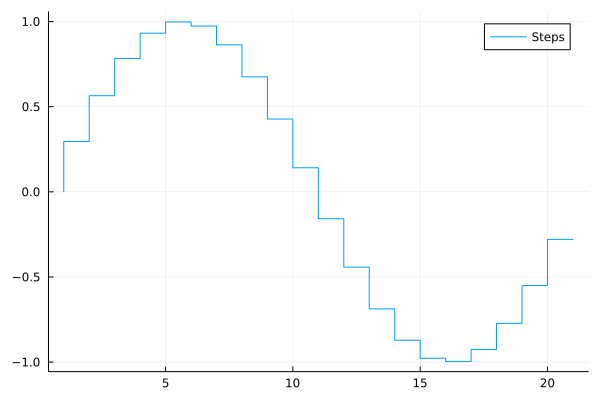
plot(sin.(0:0.3:2pi), line=:stem, marker=:star, markersize=20, ylims=(-1.1, 1.1), label="Stars")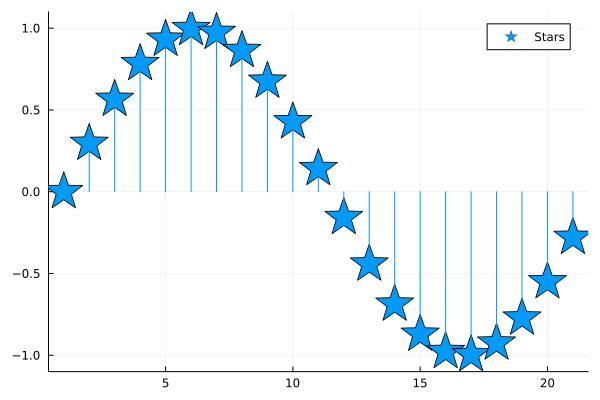
using Plots
data = rand(100, 4)100×4 Matrix{Float64}:
0.709981 0.973362 0.516176 0.363075
0.761391 0.388399 0.193247 0.724163
0.75949 0.815402 0.0369112 0.137228
0.761702 0.902961 0.330324 0.455833
0.795277 0.416984 0.945352 0.176323
0.586307 0.286339 0.499248 0.0763004
0.276137 0.765012 0.594053 0.294608
0.84172 0.687983 0.841599 0.620266
0.847136 0.393362 0.305451 0.216803
0.380462 0.687962 0.794865 0.127923
⋮
0.378577 0.106089 0.691953 0.777491
0.0571538 0.977407 0.276459 0.607869
0.674539 0.610381 0.236658 0.662346
0.278811 0.179141 0.604079 0.515615
0.695049 0.514618 0.438862 0.922791
0.924816 0.210846 0.912419 0.860384
0.508744 0.798775 0.132198 0.429291
0.5701 0.786537 0.709826 0.66473
0.83062 0.233978 0.425281 0.375258create a 2x2 grid, and map each of the 4 series to one of the subplots
plot(data, layout = 4)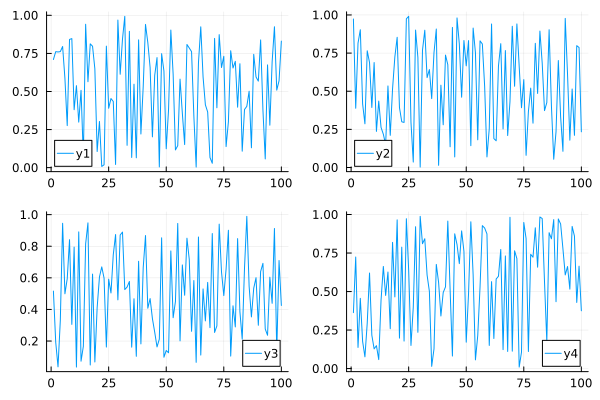
More complex grid layouts can be created with the grid(...) constructor:
plot(data, layout = grid(4, 1, heights=[0.1 ,0.4, 0.4, 0.1]))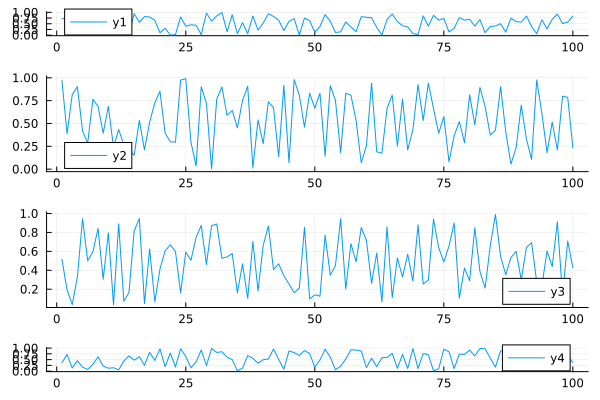
Adding titles and labels
plot(data, layout = 4, label=["a" "b" "c" "d"], title=["1" "2" "3" "4"])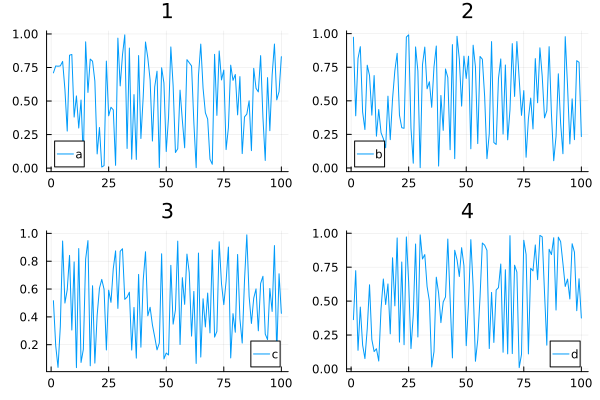
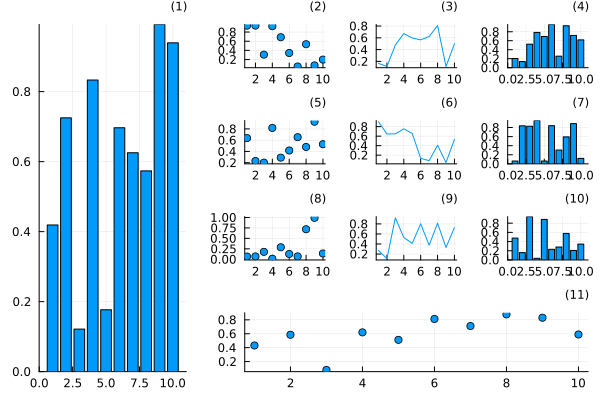
Use @layout macro
l = @layout [
a{0.3w} [grid(3,3)
b{0.2h} ]
]
plot(
rand(10, 11),
layout = l, legend = false, seriestype = [:bar :scatter :path],
title = ["($i)" for j in 1:1, i in 1:11], titleloc = :right, titlefont = font(8)
)
Use _ to ignore a spot in the layout
plot((plot() for i in 1:7)..., layout=@layout([_ ° _; ° ° °; ° ° °]))
Build subplots one by one¶
p1 = plot(sin, 0, 2pi, xlabel="x1")
p2 = plot(cos, 0, 2pi, xlabel="x2")
p3 = histogram(randn(1000), xlabel="x3")
p4 = plot(x->exp(-x^2), -3, 3, xlabel="x4")
plot(p1, p2, p3, p4)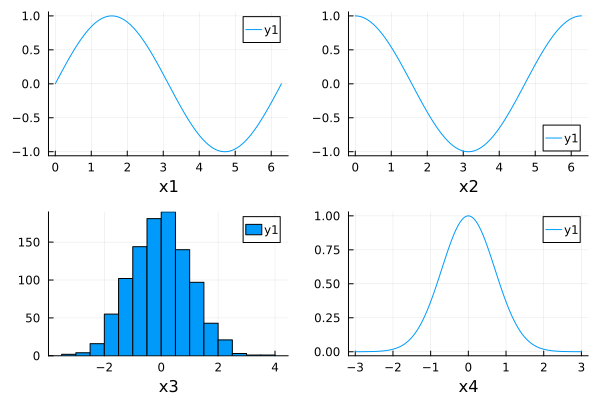
Surface plots¶
surface(x, y, z)surface(x, y, (x,y)->z)plot(x, y, z, linetype=:surface)plot(x, y, z, linetype=:wireframe)
using Plots
x = y = -10:10
f = (x , y) -> x^2 + y^2
surface(x, y, f)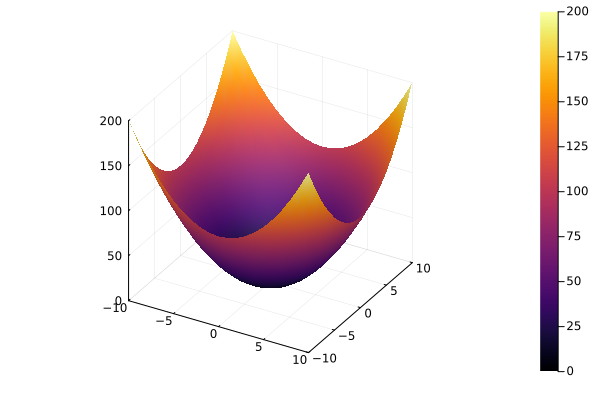
surface style
plot(x, y, f, linetype=:surface)
wireframe style
plot(x, y, f, linetype=:wireframe)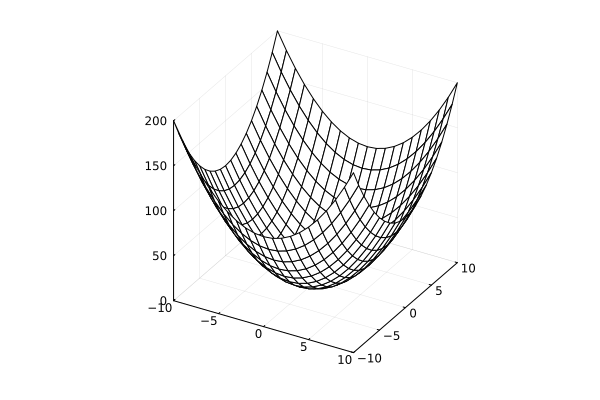
Twin Y Axis¶
plot!(twinx())
using Plots
plot(randn(100), ylabel="y1", leg=:topright)
plot!(
twinx(), randn(100)*10,
c=:red,
ylabel="y2",
leg=:bottomright,
size=(600, 400)
)
plot!(right_margin=15Plots.mm)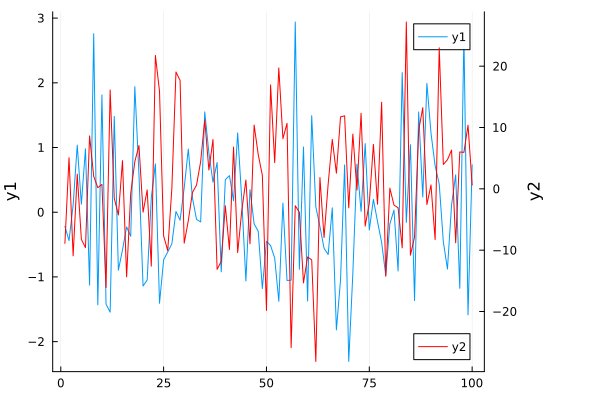
Animations¶
using Plotsdefine the Lorenz attractor
Base.@kwdef mutable struct Lorenz
dt::Float64 = 0.02
σ::Float64 = 10
ρ::Float64 = 28
β::Float64 = 8/3
x::Float64 = 1
y::Float64 = 1
z::Float64 = 1
end
function step!(l::Lorenz)
dx = l.σ * (l.y - l.x)
dy = l.x * (l.ρ - l.z) - l.y
dz = l.x * l.y - l.β * l.z
l.x += l.dt * dx
l.y += l.dt * dy
l.z += l.dt * dz
end
attractor = Lorenz()Main.var"##472".Lorenz(0.02, 10.0, 28.0, 2.6666666666666665, 1.0, 1.0, 1.0)initialize a 3D plot with 1 empty series
plt = plot3d(
1,
xlim = (-30, 30),
ylim = (-30, 30),
zlim = (0, 60),
title = "Lorenz Attractor",
marker = 2,
)
plt
pushing new points to the plot
anim = @animate for i=1:1500
step!(attractor)
push!(plt, attractor.x, attractor.y, attractor.z)
end
mp4(anim, fps = 15)[ Info: Saved animation to /tmp/jl_hABiHOJE3G.mp4
Output¶
https://
savefig("plot.pdf")The PDF format uses vector graphic format and conserves details.
Figure size e.g.
size=(750,750)affects relative fonts sizes.One can convert to other formats later using
pdftoppmorimagemagick.
Render Images¶
# using Pkg; Pkg.add("Images")
using Images
img1 = load("dog.jpg")
plot(img1)This notebook was generated using Literate.jl.